Welcome to Molecular Dynamics Analysis for IPython (ipyMD)!¶
This package aims to provide a means of producing reusable analysis of Molecular Dynamics (MD) output in the IPython Notebook.
There are many programs for 3D visualisation of MD output (my favourite being Ovito). However, there lacks a means to produce a more thorough, documented analysis of the data. IPython Notebooks are ideal for this type of analysis and so the objective of ipymd is to produce a Python package that can be used in conjuction with programmes like Ovito, to produce documented and reuseable analysis.
The aim of ipymd is to produce IPython Notebooks that include:
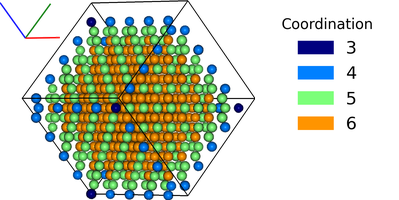
- Static images of the simulations
- Analysis of simulation data, including graphical plots
It has been created with the goal to be:
- Easy to use
- Easy to extend
chemlab It builds primarily on the chemlab package, that is an API layer on top of OpenGL. Data is parsed in standard formats, such as [pandas](http://pandas.pydata.org/) dataframes, which are easy to create and use independantly from this package, in order to extend its functionality.
| Author | Chris Sewell |
| Project Page | https://github.com/chrisjsewell/ipymd |